Next-Generation Sequencing Library Preparation
What is Next-Generation Sequencing (NGS)?
Chain termination sequencing, or Sanger sequencing, revolutionized biological research, allowing researchers to determine genome sequences for the first time. Next-generation sequencing (NGS) refers to high-throughput, massively parallel sequencing technologies for studying genetic variation associated with disease or other biological research.
These technologies are categorized by the sequencing read length – either short-read or long-read – and this length corresponds directly with the reagents used on an NGS instrument. Short-read sequencing technologies include: sequencing by synthesis, ion semiconductor, sequencing by ligation, and pyrosequencing. In contrast, NGS technologies that produce long sequencing reads are composed of single-molecule real-time sequencing and nanopore sequencing.
Assessing the quality of NGS libraries
Since the introduction of NGS platforms, DNA sequencing has rapidly evolved. However, successful sequencing depends on the development and use of high-quality NGS libraries – a critical preparation step in the NGS workflow. A typical NGS library is composed of target DNA sheared to an average size, that has undergone further enzymatic treatment specific to the library preparation kit and the sequencing platform. Numerous types of NGS library prep kits are available for different sequencing platforms and applications.
NGS workflows, similar to any genetic analysis applications, require starting your experiment with high-quality DNA and checking sample integrity at specific points throughout the workflow. For example, checking sample integrity after library preparation and before NGS analysis will enhance confidence in the results generated.
NGS library quality can be assessed by ensuring pooled library fragments are of an expected size, and that the library concentration is adequate to generate robust results. For labs attempting to assess the quality of determinants in this way, traditional hands-on methods such as agarose gel electrophoresis can be time and labor intensive. However, Agilent NGS solutions facilitate a walkway to automation and offer both high precision and sensitivity.
Delivering accurate, reproducible results
Regardless of the NGS technology used, all require a reliable nucleic acid QC process to help ensure accurate and reproducible results. Agilent has compiled several instruments and various reagent kits into an extended Automated Electrophoresis portfolio, designed to meet the evolving demands of NGS. While these instruments, including the Bioanalyzer, Fragment Analyzer, TapeStation, and Femto Pulse systems, are each designed for the QC of nucleic acids, they each offer different advantages in terms of throughput, sensitivity, speed, and resolution. Whether the demands originate from the NGS platform or from the sample type, Agilent has the QC solution.
Short-Read Sequencing
Short-read sequencing platforms, as the name suggests, use NGS libraries with a small average size, generally between 150 to 600 bp. They are widely used for the diagnostics of small mutations.
Known to provide deep coverage, short-read sequencing has been utilized in the de novo assembly of the genomes of countless organisms. Successful sequencing, however, demands optimal loading of flow cells, in turn requiring accurate determination of average library size and concentration, to perform molarity calculations.
The Agilent Automated Electrophoresis instruments help ensure successful sequencing with confident quantification, qualification, and sizing of short-read NGS library smears. For example:
- • Deliver precise analytics evaluation of various sample types with the Bioanalyzer system which is compatible with several kits covering different sizing ranges from 25 to 12,000 bp.
- The Fragment Analyzer systems offer many kits, three of which are specific for short-read libraries with a sizing range from 1 to 6,000 bp.
- From low to high throughput, the TapeStation systems provide fast run times with sizing up to 1,000 or 5,000 bp depending on the assay.
- The Femto Pulse system is ideal for situations where the short-read NGS library concentration is low or sample conservation is crucial.
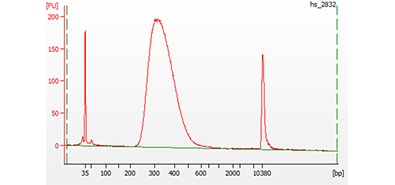
2100 Bioanalyzer system
A short-read sequencing library analyzed on a 2100 Bioanalyzer system with a High Sensitivity DNA kit.
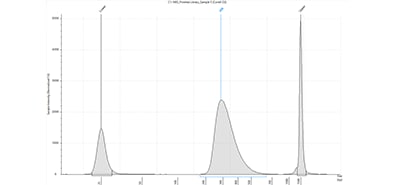
TapeStation system
Final library of a SureSelectXT workflow analyzed with the D1000 ScreenTape assay using a TapeStation system.
Short-read sequencing provides many sub applications to address challenging sample types and unique questions. Different types of short-read sequencing libraries can enable identification of epigenetic regions, diminish bias in sequencing libraries with PCR-free libraries, and deepen understanding of gene expression.
ChiP-Seq
Chromatin Immunoprecipitation sequencing (ChIP-seq) provides crucial information on protein-DNA interactions, applying NGS to regions of genomic DNA involved with protein binding. Typical ChIP-seq workflows are resource-intensive, requiring high concentration samples for crucial QC checkpoints during library construction, limiting the application of ChIP-seq to abundant cell types.
The sensitivity of the Automated Electrophoresis instruments alleviates this restriction, dramatically decreasing the amount of sample required for quality control analysis. Successfully quantify ChIP-seq libraries with:
- The High Sensitivity DNA kit for the 2100 Bioanalyzer system
- The HS NGS Fragment Kit with the Fragment Analyzer systems
- The D1000 assay for the TapeStation systems
TapeStation system
Final library of a ChIP-Seq workflow analyzed on a TapeStation system with the D1000 ScreenTape assay. Successful libraries show a narrow library distribution with a peak size of 120–130 bp larger than the starting material. In this example, the final library shows a maximum peak size of 318 bp, correlating to an insert size of 200 bp.
PCR-Free Libraries
Many NGS library construction protocols require at least one PCR amplification step to ensure that there is enough material to construct a library at a sufficient concentration. While valid, amplification steps frequently introduce bias and error into the sequencing libraries. PCR-free libraries eliminate amplification steps, reducing error and removing bias from NGS library construction. Problematically, PCR-free libraries are often at such a low concentration that legacy methods of NGS library QC are unable to size, quantify, and qualify the libraries.
The Femto Pulse system provides the necessary sensitivity for the evaluation of PCR-free libraries. The kit enables the fast and accurate QC analysis of PCR-free libraries, providing researchers with the results they need when they need them.
Small RNA Libraries
Small RNAs are involved in many important biological processes, notably gene expression. Small RNA sequencing provides researchers with further insight into these processes. Sequencing libraries made from small RNA present novel QC challenges due to the small smear size and narrow smear range.
The precision resolution of the 2100 Bioanalyzer and the Fragment Analyzer systems alleviate issues with the accurate analysis of small RNA NGS libraries. In the example shown below, with the HS Small Fragment kit for the Fragment Analyzer systems, researchers can accurately size, quantify, and qualify small RNA NGS libraries with ease.
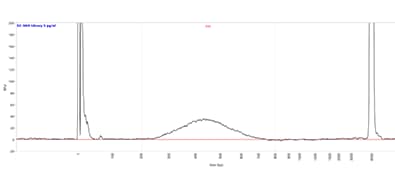
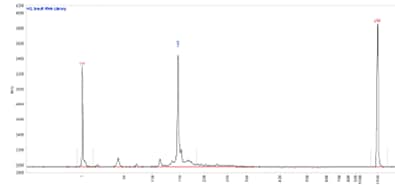
Fragment Analyzer system
Small RNA library separated using the HS Small Fragment kit. Capillary electrophoresis performed on a Fragment Analyzer equipped with a short capillary array, 33 cm.
Recommended assays for short-read sequencing libraries
| 2100 Bioanalyzer system | Fragment Analyzer system | TapeStation system | Femto Pulse system |
|
High Sensitivity DNA kit (p/n 5067–4626) |
NGS Fragment kit (p/n DNF-473) |
D1000 ScreenTape assay (p/n 5067–5582 and 5067-5583) |
Ultra Sensitivity NGS kit (p/n FP-1101) |
| May also use: |
HS NGS Fragment kit (p/n DNF-474) |
D5000 ScreenTape assay (p/n 5067–5588 and 5067-5589) |
|
|
DNA 1000 kit (p/n 5067–1504) |
HS Small Fragment kit (DNF-477) |
High Sensitivity D1000 ScreenTape assay (p/n 5067–5584 and 5067-5585) |
|
|
DNA 7500 kit (p/n 5067–1506) |
High Sensitivity D5000 ScreenTape assay (p/n 5067–5592 and 5067-5593) |
||
|
DNA 12000 kit (p/n 5067–1508) |
For Research Use Only. Not for use in diagnostic procedures.
Long-Read Sequencing
Long-read sequencing libraries are composed of fragments ranging from 20 to 40 kb, and offer great value and utility when accessing genetic information. By sequencing these long fragments of DNA enables researchers with deeper insight into genome architecture.
Agilent’s sample QC solutions enable researchers to assess large DNA fragments, including input genomic DNA and long-read sequencing libraries, to help ensure robust NGS results.
- • Quantify, qualify, and size DNA and RNA samples with the Femto Pulse system. The Genomic DNA 165 kb kit (p/n FP-1002) provides optimized pulsed-field separation methods for enhanced resolution of samples up 165 kb.
- • Carry out reliable QC of nucleic acids with either one of two reagent kits for the Fragment Analyzer systems are available: the Large Fragment kit (p/n DNF-492) provides reliable sizing through 20 kb and the HS Large Fragment 50 kb kit (p/n DNF-464)Provides accurate sizing and quantification of large DNA fragments through 60 kb.
- The TapeStation systems provide fast analysis of samples up to 60,000 bp in less than 2 minutes per sample with the Genomic DNA ScreenTape assay (p/n 5067–5365 and 5067-5366).
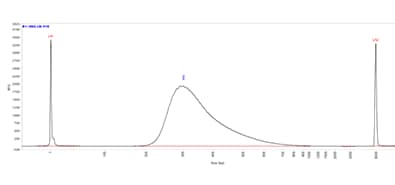
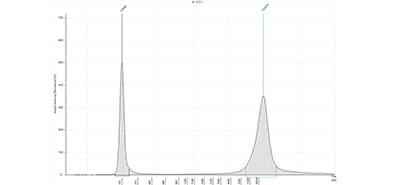
Femto Pulse system
Long-read sequencing library separated on the Femto Pulse system using the Genomic DNA 165 kb kit (FP-1002).
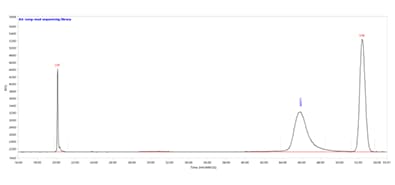
Fragment Analyzer system
Long-read sequencing library separated using the HS Large Fragment 50 kb kit. Capillary electrophoresis was performed on a Fragment Analyzer equipped with a short capillary array, 33 cm.
TapeStation system
Electropherogram of reference gDNA analyzed on a TapeStation system using the DNA ScreenTape assay. With a DIN of 9.7, the sample qualifies as starting material for DNA library preparation workflows.
For Research Use Only. Not for use in diagnostic procedures.
Solutions for accurate, versatile analysis of DNA samples
Of the methods for nucleic acid quality control analysis, the Agilent automated electrophoresis instruments provide accurate and versatile analysis of DNA samples. The Bioanalyzer, Fragment Analyzer, TapeStation, and Femto Pulse systems provide variable throughput, high sensitivity detection, and broad sizing ranges for reliable nucleic acid QC in NGS workflows of any type.
| Instrument | Part Number(s) | Reagent Kit Name | Sizing Range | Quantification Range* |
|---|---|---|---|---|
| 2100 Bioanalyzer system | 5067–4626 | High Sensitivity DNA kit | 50 to 7,000 bp |
Fragment: 5 to 500 pg/µL Smear: 0.1 to 10 ng/µL |
| 5067–1504 | DNA 1000 kit | 25 to 1,000 bp | 0.5 to 50 ng/µL | |
| 5067–1506 | DNA 7500 kit | 100 to 7,500 bp | 0.5 to 50 ng/µL | |
| 5067–1508 | DNA 12000 kit | 100 to 12,000 bp | 0.5 to 50 ng/µL | |
| Fragment Analyzer systems |
DNF-473-0500 DNF-473-1000 |
NGS Fragment kit | 100 to 6,000 bp |
Fragment: 0.1 to 10 ng/µL Smear: 5 to 100 ng/µL |
|
DNF-474-0500 DNF-474-1000 |
HS NGS Fragment kit | 100 to 6,000 bp |
Fragment: 5 to 500 pg/µL Smear: 50 to 5,000 pg/µL |
|
|
DNF-477-0500 DNF-477-1000 |
HS Small Fragment kit | 50 to 1,500 bp |
Fragment: 5 to 500 pg/µL> Smear: 100 to 5,000 pg/µL |
|
| DNF-464-0500 | HS Large Fragment 50 kb kit | 75 to 50,000 bp |
Fragment: 5 to 600 pg/µL Smear: 50 pg/µL to 5 ng/µL |
|
|
DNF-492-0500 DNF-492-1000 |
Large Fragment kit | 50 to 20,000 bp |
Fragment: 0.1 to 10 ng/µL Smear: 5 to 100 ng/µL |
|
| TapeStation systems |
5067-5582 (ScreenTape) 5067-5583 (Reagents) |
D1000 ScreenTape | 35 to 1,000 bp |
Fragment: 0.1 to 50 ng/µL Smear: 5 to 100 ng/µL |
|
5067-5584 (ScreenTape) 5067-5585 (Reagents) |
High Sensitivity D1000 ScreenTape | 35 to 1,000 bp |
Fragment: 10 to 1,000 pg/µL Smear: 0.5 to 15 ng/µL |
|
|
5067-5588 (ScreenTape) 5067-5589 (Reagents) |
D5000 ScreenTape | 100 to 5,000 bp | 0.1 to 50 ng/µL | |
|
5067-5592 (ScreenTape) 5067-5593 (Reagents) |
High Sensitivity D5000 ScreenTape | 100 to 5,000 bp | 10 to 1,000 pg/µL | |
|
5067-5565 (ScreenTape) 5067-5566 (Reagents) |
Genomic DNA ScreenTape | 200 to >60,000 bp | 10 to 100 ng/µL | |
| Femto Pulse system | FP-1101-0275 | Ultra Sensitivity NGS kit | 100 to 6,000 bp |
Fragment: 0.05 to 5 pg/µL Smear: 5 to 250 pg/µL |
| FP-1002-0275 | Genomic DNA 165 kb kit | 1.3 to 165 kb |
Fragment: 0.3 to 30 pg/µL Smear: 5 to 500 pg/µL |
For Research Use Only. Not for use in diagnostic procedures.
NGS Application Notes
Comparison of the Agilent HS Small Fragment Kit and the Agilent HS NGS Fragment Kit on the Agilent 5200 Fragment Analyzer SystemThe Agilent Fragment Analyzer equipped with the short or ultra-short array offers consistent sizing and quantification of gDNA smears.
The Agilent 4150 TapeStation system performs reliable sample quality control (QC) through various steps of the SureSelect XT HS target enrichment and nextgeneration sequencing (NGS) library preparation workflow on the Agilent Magnis NGS Prep system.
Evaluation of the Agilent 4150 TapeStation system with ScreenTape assay to monitor exome and transcriptome library preparation for systems biology omics analysis.
This application note addresses and demonstrates the utility of two automated electrophoretic platforms - TapeStation and Bioanalyzer systems - at various QC check-points throughout the Agilent NGS product portfolio
This Application Note describes quality control (QC) steps in various Illumina next-generation sequencing (NGS) workflows provided as a service by the German Cancer Research Center (DKFZ) Genomics and Proteomics Core Facility.
For Research Use Only. Not for use in diagnostic procedures.