QuikChange II Site-Directed Mutagenesis Kits - Details & Specifications
The QuikChange II site-directed mutagenesis kit is used to make point mutations, replace amino acids, and delete or insert single or multiple adjacent amino acids. The QuikChange II site-directed mutagenesis method is performed using PfuUltra high-fidelity (HF) DNA polymerase for mutagenic primer-directed replication of both plasmid strands with the highest fidelity. The basic procedure utilizes a supercoiled double-stranded DNA (dsDNA) vector with an insert of interest and two synthetic oligonucleotide primers, both containing the desired mutation (see Figure 1). The oligonucleotide primers, each complementary to opposite strands of the vector, are extended during temperature cycling by PfuUltra HF DNA polymerase, without primer displacement. Extension of the oligonucleotide primers generates a mutated plasmid containing staggered nicks. Following temperature cycling, the product is treated with Dpn I. The Dpn I endonuclease (target sequence: 5´-Gm6ATC-3´) is specific for methylated and hemimethylated DNA and is used to digest the parental DNA template and to select for mutation-containing synthesized DNA.6 (DNA isolated from almost all E. coli strains is dam methylated and therefore susceptible to Dpn I digestion.) The nicked vector DNA containing the desired mutations is then transformed into competent cells.
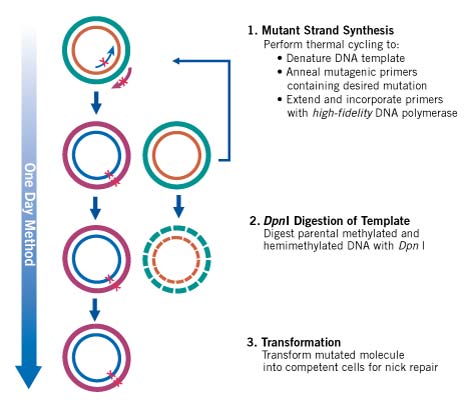
Figure 1. Overview of the QuikChange II Site-directed Mutagenesis Method.
QuikChange II Site-Directed Mutagenesis Kit
For short (4kb – 8kb) targets. Includes XL1-Blue Competent Cells.
QuikChange II XL Site-Directed Mutagenesis Kit
For long (8 kb – 14kb) or difficult targets. Includes XL10-Gold Ultracompetent Cells.
QuikChange II-E Site-Directed Mutagenesis Kit
For researchers performing mutagenesis via transformation into electroporation-competent cells. Includes XL1-Blue Electroporation-competent Cells.
For Research Use Only. Not for use in diagnostic procedures