Genomic DNA Quality Control Analysis
Genomic DNA Quality Control Analysis
First isolated in the mid-1800s, genomic DNA (gDNA) comprises of the chromosomal DNA of an organism. It contains all an organism’s biological information to direct its cellular function, which is also passed on to the next generation.
The gDNA sequence carries information organized in genes and other discrete genetic elements, which researchers can access by isolating genetic material and sequencing or manipulating it. This access is regulated by the genome’s organization or structure, and chemical and physical modifications to the DNA, including the associated binding of small molecules and proteins. Genomic analysis helps scientists better understand the functions, interactions, and regulation of specific genes that control the development of an organism.
Whether your research application uses next-generation sequencing (NGS), gene editing, cloning, or genotyping, beginning your experiments with high-quality, high-integrity DNA helps ensure robust results.
Agilent offers a portfolio of solutions for gDNA quality analysis, including instruments, reagent kits, and software to enable you to successfully assess DNA size, integrity, and concentration. Achieve reliable results with automation and scalable throughput.
Genomic DNA Quality Control Analysis you can rely on
Molecular biologists and geneticists employ many applications that require gDNA and high-quality DNA samples to generate reliable, robust results. Assessment of sample quality is important.
The Agilent Automated Electrophoresis portfolio includes a range of instruments that offer advantages in terms of throughput scalability, sensitivity, speed, and resolution. Read on to find out how you can conduct superior assessment of intact gDNA, including accurate qualification, sizing, and quantification.
Increase Throughput with Fragment Analyzer Systems
Agilent’s Fragment Analyzer systems provide you with robust gDNA analysis solutions to fit any throughput need – from 12 samples in about one hour, to over 2,400 samples in 24 hours.
There are two kits designed to analyze gDNA:
- The Genomic DNA 50 kb kit (p/n DNF-467) provides confident assessment of gDNA smears through 60 kb. A single 200-fold dilution greatly simplifies sample preparation, conserving researcher time.
- The HS Genomic DNA 50 kb kit (p/n DNF-468) offers sizing of gDNA smears through 60 kb, for lower concentration samples.
GQN: An Empirical Assessment of Quality
Agilent’s fragment analysis solutions allow for easy and empirical assessment of gDNA quality with a dynamic quality metric. ProSize data analysis software calculates a Genomic Quality Number (GQN) for the gDNA smear as a value between 0 and 10, simplifying comparison of quality.
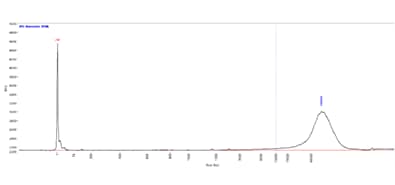
The image shows the results when gDNA separated using the Genomic DNA 50 kb kit on a Fragment Analyzer system equipped with a short capillary array. The GQN was applied to this sample with a size threshold of 10,000 bp (purple line) and GQN was calculated at 9.0, indicating high-quality gDNA.
Smear Analysis To Improve Accuracy And Precision
Agilent’s fragment analysis solutions also support specific analysis of concentration and size (bp) of gDNA smears within a user-defined sizing range. The Smear Analysis function provides researchers with the accuracy and precision demanded by downstream applications.
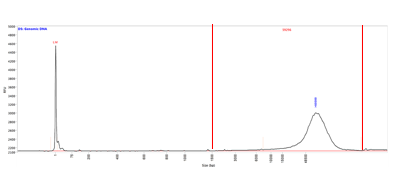
In this smear analysis, the smear range is represented by the dotted red lines. The size of the highest peak within the smear is indicated by the blue number over the peak. In contrast, the Smear Analysis determines an average smear size, shown by the red number on the top of the electropherogram and in the inset table.
Simplified Electrophoresis with Agilent TapeStation systems
High-quality input gDNA samples are essential for downstream applications, such as NGS.
The TapeStation systems offer easy and fast automated electrophoretic separations of genomic DNA. This is supplemented by the Genomic DNA ScreenTape assay (p/n 5067-5365 and 5067-5366), which provides accurate quantitation and reliable quality analysis of up to 96 samples from 200 through 60,000 bp in less than two minutes per sample.
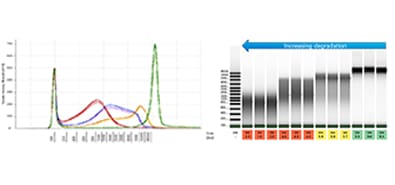
The TapeStation software provides an an automated and objective quality score, the DNA integrity number (DIN). The DIN ranges from 1 to 10 and indicates the quality of a gDNA sample by assessing its distribution across the entire sizing range. A lower DIN correlates to more degraded sample, while a high DIN would indicate a good quality sample, allowing researchers to make more informed decisions.
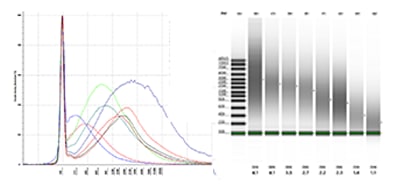
gDNA samples at four degradation stages were analyzed on a TapeStation system with the Genomic DNA ScreenTape assay to determine DNA integrity and sample concentration. The image above shows (A) the gel image with corresponding DIN displayed below each lane, and (B) the corresponding electropherogram overlay of all the gDNA samples.
Detailed information and accurate sizing of gDNA
For applications that require more detailed information and accurate DNA sizing above 60 kb, such as long read sequencing, the Femto Pulse system is ideal. A meticulously designed optical detection platform with unprecedented sensitivity, the Femto Pulse system is flexible enough to handle any workflow scenarios. The Genomic DNA 165 kb kit (p/n FP-1002) uses pulsed-field capillary electrophoresis, to size high molecular weight gDNA smears through an unprecedented 165 kb, in about 1.5 hours.
Additionally, the ProSize data analysis software tools used with the Fragment Analyzer systems can be used with the large gDNA smears analyzed on the Femto Pulse system. This provides you with even more detailed information about high molecular weight samples.
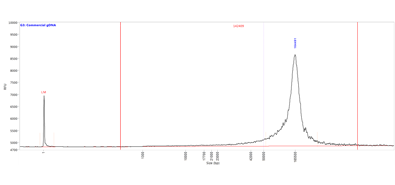
The image shows the results when pulsed-field capillary electrophoresis was performed using a Femto Pulse system equipped with a Femto Pulse 12-Capillary Array. The dotted blue line on the electropherogram represents the size threshold selected for GQN determination, and the vertical red lines denote the smear analysis range. For this sample, with a user-defined threshold of 50,000 bp, the GQN is 8.6, indicating that 86% of the sample is larger than the threshold.
For Research Use Only. Not for use in diagnostic procedures.
Sensitive Analysis of Degraded and FFPE gDNA
gDNA that has been fragmented or degraded – for instance, formalin-fixed, paraffin-embedded (FFPE) gDNA – presents unique challenges for QC analysis. Quality scoring of FFPE gDNA and degraded gDNA is complex, often requiring researchers to make a best guess when using conventional agarose gel electrophoresis.
For instance, gDNA extracted from FFPE tissue often undergoes shearing and cross-linking with proteins and other molecules, frequently resulting in a complex and convoluted separation profile. Agilent’s range of automated electrophoresis systems, including the Fragment Analyzer, TapeStation, and Femto Pulse systems enable researchers to extract reliable results even from FFPE gDNA.
While FFPE gDNA generally has a lower integrity, knowing the quality of the sample can allow a user to modify certain workflows for these samples. For example, fragmentation conditions could be adapted based on a sample’s initial integrity. Agilent’s fragment analysis and Femto Pulse solutions allow for accurate assessment of FFPE gDNA quality and integrity with two metrics: the Genomic Quality Number, and the DNA integrity number.
The ProSize data analysis software calculates a Genomic Quality Number (GQN) for the gDNA smear as a value between 0 and 10, enabling an empirical quality scoring strategy. The Genomic DNA ScreenTape assay for the TapeStation systems provides an objective measurement of DNA integrity through the DNA integrity number (DIN).
Calculating the GQN of a sample of FFPE gDNA
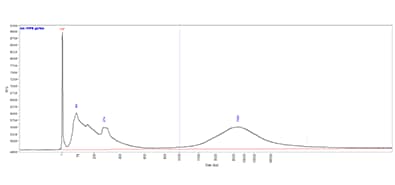
The image shows the results of FFPE gDNA separated using the Genomic DNA 50 kb kit on a Fragment Analyzer system equipped with a short capillary array. A size threshold of 1,000 bp, denoted by the dotted blue line, provided a GQN of 3.7.
Working out the DIN of a sample of FFPE gDNA
The image shows the results of FFPE gDNA samples of varying integrity separated on a TapeStation system using the Genomic DNA ScreenTape assay. The results are shown as both an electropherogram overlay and a digital gel image, with the DIN listed below each gel lane.
For Research Use Only. Not for use in diagnostic procedures.
Discover Agilent’s comprehensive portfolio of Genomic DNA (gDNA) Analysis Kits
Agilent offers several instrument-based solutions for the effective and reliable analysis of gDNA. The Fragment Analyzer, TapeStation, and Femto Pulse systems, with the appropriate assays and reagent kits, provide superior analysis with unmatched resolution and sizing capabilities.
| Instrument | Part Number | Reagent Kit Name | Sizing Range | Quantitative Range | Quality Metric |
|---|---|---|---|---|---|
| Fragment Analyzer systems | DNF-467-0500 | Genomic DNA 50 kb kit | 75 to 60,000 bp | 25 to 250 ng/µL | GQN |
| DNF-468-0500 | HS Genomic DNA 50 kb kit | 75 to 60,000 bp | 0.3 to 12 ng/µL | GQN | |
| TapeStation systems |
5067–5365 5067-5366 |
Genomic DNA ScreenTape Genomic DNA reagents |
200 to >60,000 bp | 10 to 100 ng/µL | DIN |
| Femto Pulse system | FP-1002-0275 | Genomic DNA 165 kb kit | 1,300 to 165,000 bp |
Fragment: 5 to 500 pg/µL Smear: 0.3 to 30 pg/µL |
GQN |
For Research Use Only. Not for use in diagnostic procedures.
Read more about gDNA analysis in our application notes.
Assessment of Genomic DNA Quality with the Agilent 5200 Fragment Analyzer SystemThe Agilent 5200 Fragment Analyzer offers effortless analysis of genomic DNA (gDNA) with the Genomic DNA 50 kb Kit and the HS Genomic DNA 50 kb Kit.
Separation of HMW gDNA, providing accurate, reproducible sizing and quality control using the Agilent Femto Pulse System.
The Agilent Femto Pulse system is the only automated system capable of replacing overnight PFGE analysis of HMW gDNA in only 70 minutes, saving time and money.
The Agilent Femto Pulse System is the only instrument capable of replacing pulsed-field gel electrophoresis for the analysis of high molecular weight (HMW) gDNA.
QC steps including quantification and qualification of the DNA, and can be done with the 4200 TapeStation System.
This Application Note demonstrates the use of the Agilent 4150 TapeStation system as a quality control (QC) tool to analyze DNA samples stored at the Heidelberg CardioBiobank (HCB).
For Research Use Only. Not for use in diagnostic procedures.