Access Agilent eNewsletter November 2015
>> Update My Profile | Subscribe to Access Agilent | Article Directory

Using untargeted mass spectrometry for pesticide detection in food
By John Lee
Agilent Food Market Manager
As the food supply chain expands and becomes increasingly global there is increasing diversity in the types of pesticide contamination. For example, there are over 1,000 approved compounds that farmers might apply above the prescribed amount. Banned compounds cannot be discounted as potential risks, either. Therefore, choosing which pesticides to test for protection of public health is a growing challenge, and so many government labs are looking at untargeted mass spectrometry (MS) techniques to collect these key data.
In this first article in a two-part series on pesticide monitoring, we will explore how mass spectrometers that operate in full-scan mode can be effective tools for pesticide detection and measurement. This approach places no limits on the range of pesticides that a laboratory can consider when subsequently interrogating the data. Furthermore, new compounds not previously considered in a surveillance scheme can be easily added, and searched for in both new and previous data. Matrix effects are of increasing concern with full scan especially given the number of pesticides being monitored and low detection limits. This is why sample preparation is increasingly important.
 Enlarge
Enlarge
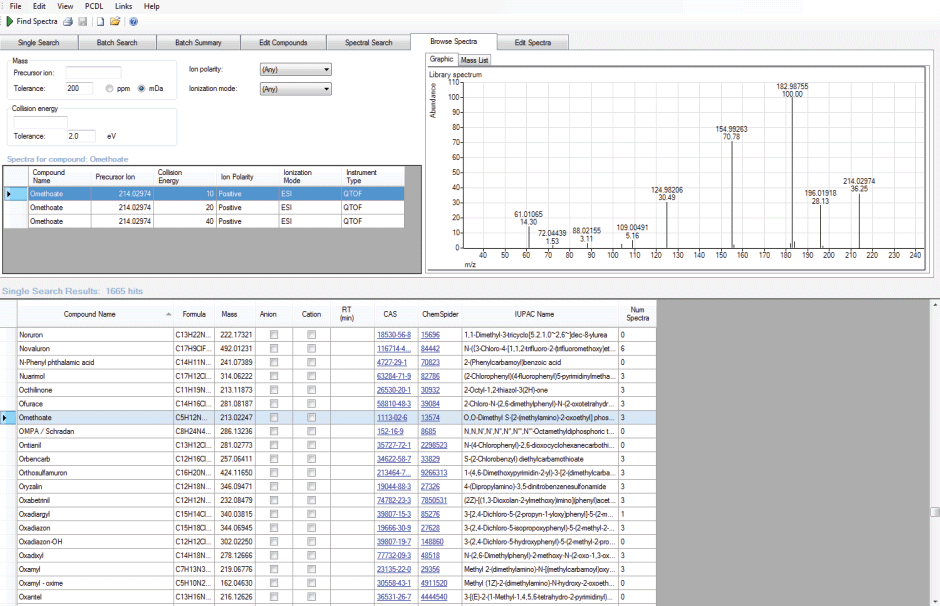
Figure 1. PCDL software showing the pesticide library and the exact mass spectrum of omethoate.
 Enlarge
Enlarge
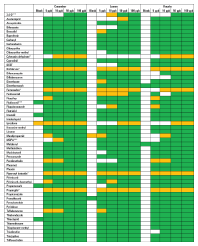
Table 1. Results of screening and verifying pesticides in fruits and vegetables by target MS/MS acquisition and library searching.
 Enlarge
Enlarge
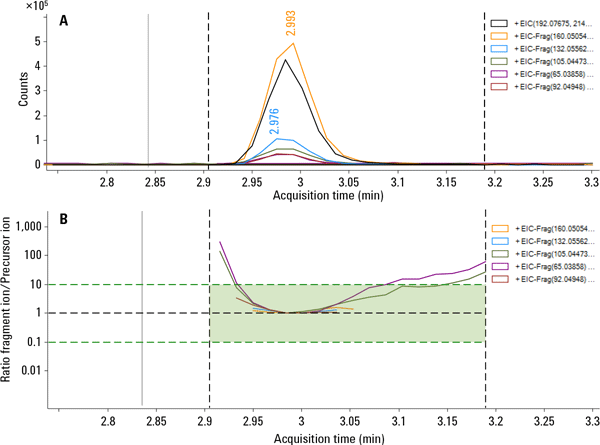
Figure 2. Overlaid ion chromatograms (A) and calculated coelution plot (B) from a pesticide screening analysis.
 Enlarge
Enlarge
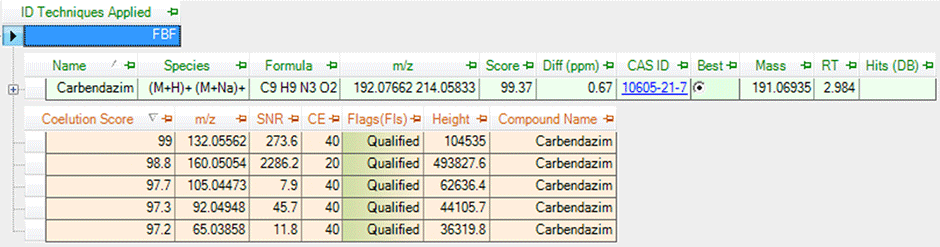
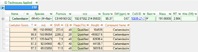
Figure 3. Software analysis results showing five valid quantifier fragments for carbendazim.
Expanding pesticide coverage requires appropriate sample preparation
Labs wishing to implement untargeted MS must find effective sample preparation strategies. Sufficient food matrix must be removed to allow sensitive and reliable pesticide detection while ensuring good recovery of the compounds themselves. Increasing the diversity of compounds is a challenge, and there will always be some compounds that will require separate treatment. However, Agilent has shown how a large proportion of pesticide can be extracted using well-designed QuEChERS protocols, even when dealing with challenging matrices like high-fat foods.
Agilent Q-TOF MS is the right tool for full scan screening
For the analysis itself, Q-TOF mass spectrometry provides excellent full-scan performance with LC and GC separation. To cover the whole range of potential pesticides, both separation modes are necessary. However, even after cleanup, matrix ions still dominate in the MS, regularly co-occurring in abundance with the pesticide ions of interest. Agilent Q-TOF instruments measure these ions with high resolution and <5ppm mass accuracy, enabling detection at levels much lower than the matrix ions around them. Additionally, Q-TOF performance is not affected by matrix ions that enter the MS at the same time as a pesticide of interest. This is not true for ion trap mass spectrometers. Unlike trapping instruments, Q-TOF sensitivity to pesticide ions remains unchanged by the population of other ions that co-occur. Hence, the instrument can always collect data in a fully independent (untargeted) manner without the need for quadrupole filtering. The full potential benefits of untargeted analysis are therefore realized with Q-TOF technology.
Global collaboration creates new exact mass pesticide library
To help realize this potential, Agilent collaborated with some key government labs around the world to build solutions and best practice for the use of Q-TOF MS for qualitative pesticide screening. In a project with the State Laboratory for Health and Veterinary Affairs in Dresden, Germany, an exact mass LC/MS/MS library for pesticides was used to drive LC/Q-TOF screening of pesticides in fruit and vegetable extracts, and to reliably verify results through MS/MS library matching to ensure very low false-positive rates (Figure 1).
As a proof of principle, the method was successfully validated for screening and identifying more than 50 pesticides in three different commodity groups (Table 1). When applied to real-world samples from a routine monitoring program, all pesticides detected previously were also identified by this approach.
In fact, the library used in this work contained 770 compounds with exact mass MS/MS spectra for both polarities and with up to three different collision energies. This library was created under carefully audited conditions and now forms part of the Agilent PCDL (Personal Compound Database and Library) portfolio. It is also part of the Agilent LC/MS pesticide application kit that includes, in total, over 1600 searchable compound entries.
Agilent LC/Q-TOF data can be searched and verified using simple data review
It is also possible to use Agilent PCDL libraries with Agilent all ions MS/MS data acquisition for LC/Q-TOF. This is a fully untargeted data collection mode, but one that can collect both molecular ion and fragment ion information at the same time. Therefore, it is unnecessary for testing labs to collect MS/MS spectra to verify their finds. Verification is instead achieved through coelution scoring (Figure 2), which tracks the elution profiles of all fragment ions in the PCDL. Data review is simpler and verifications based on fragment ions can even be made retrospectively (Figure 3).
Agilent offers its Q-TOF advantage with both LC and GC separation
With GC/Q-TOF, even more ions can be produced during an analysis – making even more important the ability of Q-TOF to monitor all of these ions equally. With truly untargeted data acquisition, processing strategies similar to those for LC/Q-TOF can then be applied. We will look at GC/Q-TOF in the next article of this series. In the meantime, we suggest that you explore the comprehensive Agilent LC/MS and GC/MS multiresidue solutions for pesticide analysis in foods.
>> Update My Profile | Subscribe to Access Agilent | Article Directory
Table 1.
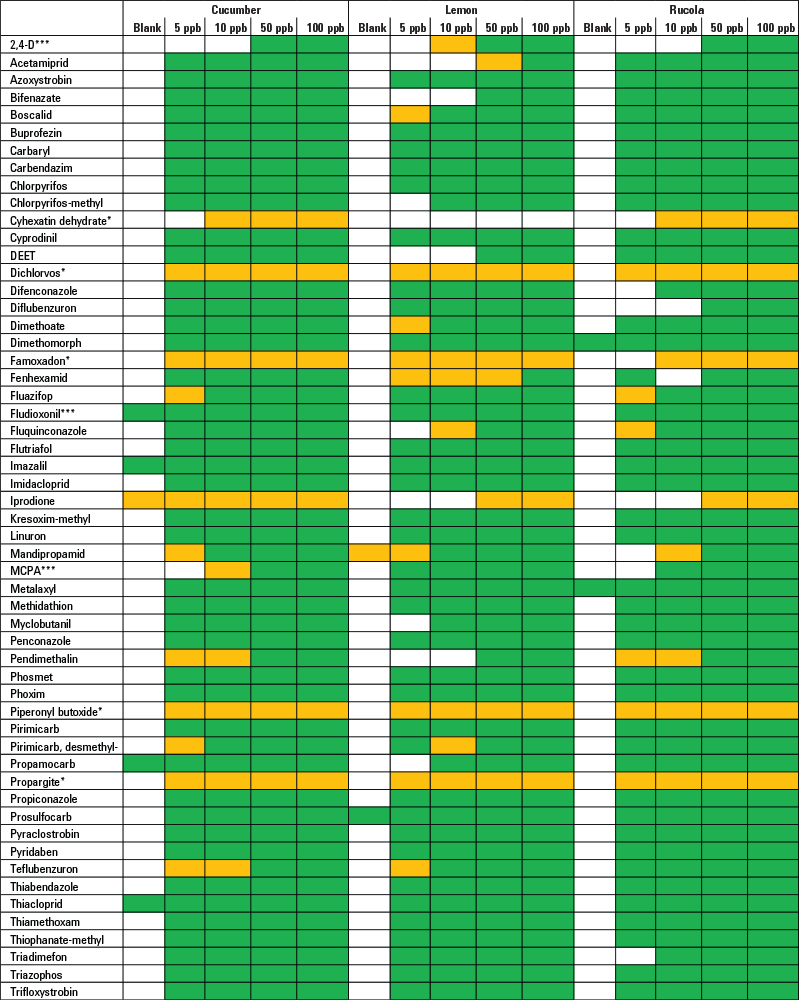
*No spectrum available for predominant compound adduct
***Compound results acquired in negative-ion mode
Results of screening and verifying pesticides in fruits and vegetables by target MS/MS acquisition and library searching (green, compound automatically found and presence verified by MS/MS library confirmation; yellow, compound automatically found but no qualified MS/MS spectrum available).