Access Agilent eNewsletter May 2015
>> Update My Profile | Subscribe to Access Agilent | Article Directory

Using Agilent Integrated Systems to understand species-specific toxicity
By Patrick D. McMullen and Melvin E. Andersen
The Hamner Institutes for Health Sciences
and Mary T. McBride
Agilent Director, Government Relations
Exposure to environmental chemicals often leads to physiological outcomes that differ between humans and rodents, calling into question the applicability of animal-based toxicity testing for human risk assessment. For example, peroxisome proliferators, including endogenous fatty acids and drugs, induce fatty acid metabolism in humans. Receptors activated by peroxisome proliferators have therefore been used as therapeutic targets for dyslipidemia (abnormal amount of lipids in the blood) and diabetes. While peroxisome proliferators regulate fatty acid metabolism in rodents, they also influence peroxisome assembly, inflammatory responses, and cell proliferation. Many peroxisome proliferators are potent carcinogens in rodents but do not appear to be carcinogenic in humans [1-3]. These differences in response epitomize the need to translate toxicological research across species and better understand the underlying molecular mechanisms. Peroxisome proliferator-activated receptors (PPARs) are nuclear receptors that function as transcription factors. In this article, we examine the results of a series of experiments designed to improve understanding of species-specific differences in PPAR-alpha (PPARα) activation at the molecular level. [4]
 Enlarge
Enlarge
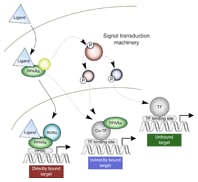
Figure 1. The canonical mechanism of PPARα-mediated response and two additional binding mechanisms suggested by our results.
Combining data streams to map PPARα networks
First, we combined high-throughput gene expression, ChIP-seq, and metabolomics experiments to characterize the response in primary hepatocytes for both humans and rodents to the PPARα-selective agonist GW7647. Gene expression was measured over multiple concentrations and times. We found a substantially larger set of differentially expressed genes (2,713) in the rat, as compared to (192) in human hepatocytes. The lipid metabolism transcription program induced by GW7647 is remarkably well conserved between the two species, but a large set of genes are down regulated in rats that is not differentially expressed in humans. These additional genes include a suite of genes that govern developmental processes, which could account for the increased toxicity of peroxisome proliferators in rodents.
The ChIP-seq experiments were performed at key time and dose points for both species to assess genomic binding of PPARα. Coupling these results with a computational motif search revealed that the canonical nuclear receptor response mechanism (that is, direct binding) accounted for only a small fraction of the detected response (24% of genes in rats, 32% in humans). This suggests a larger role for indirect PPARα transcriptional mechanisms than was previously thought. Figure 1 shows the direct binding of activated PPARα to a well-established consensus-binding site, the peroxisome proliferator response element (PPRE). However, this mechanism only accounts for about 30% of observed changes in gene expression. Our results support two additional mechanisms: indirect genomic binding and unbound nongenomic interactions.
 Enlarge
Enlarge
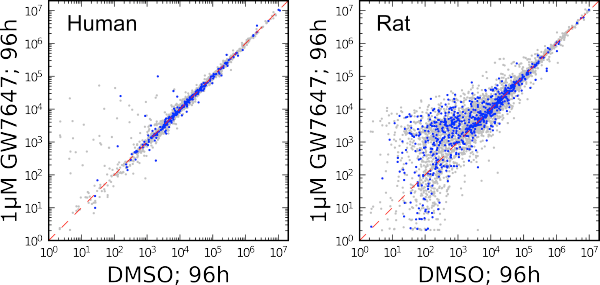
Figure 2. Comparison of metabolomic features from the secretomes of cultured primary human hepatocytes (left) and rat hepatocytes (right).
Orthogonal data enable deeper insights
To assess metabolomics changes caused by PPARα activation, we performed liquid chromatography high-resolution mass spectrometry (LC-HRMS) using nontargeted metabolomics analysis. Primary hepatocytes from both rats and humans were cultured and treated with GW7647, and the secretome analyzed across multiple time/dose points. Samples were analyzed using an Agilent 6500 Series Accurate-Mass Quadrupole Time-of-Flight (Q-TOF) LC/MS coupled to an HPLC Agilent 1290 Infinity LC via an electrospray interface (ESI). Two orthogonal UPLC separation methods, using normal-phase HILIC and Agilent ZORBAX Plus C8 reversed-phase columns, were used to maximize detection.
Activation of hepatocytes by GW7647 causes extensive alterations in fatty acid metabolism and large increases in peroxisome numbers. There were many more altered features in rat hepatocytes as compared to human (Figure 2). While additional analysis is still underway to unequivocally identify and confirm metabolites, a number of provisionally identified metabolites parse into several categories of biologically active compounds. These include varanic acid, bile acids, di- and monoglycerides containing plasmalogens [2]. In Figure 2, both cell cultures were dosed with PPARα-specific agonist GW7647 at a concentration of 1 mM, with samples analyzed 96 hours after exposure. Blue markers indicate features putatively classified as lipids in the Human Metabolome Database ontology.
Agilent integrated solutions for toxicology
Agilent provides an unrivaled suite of analytical platforms for integrated multiomics discovery, with solutions for genomics, metabolomics, and proteomics. Explore the full range of options and take advantage of an integrated systems toxicology approach with your research. Be sure to look at the latest version of our GeneSpring Bioinformatics software, designed for pathway-centric multiomic data integration.
References
- Bentley, P. et al. Food Chem. Toxicol. 31, 857-907 (1993).
- Cattley, R. et al. Regul. Toxicol. Pharmacol. 27, 47-60 (1998).
- Corton, J. C. et al. Crit. Rev. Toxicol. 44, 1-49 (2014).
- McMullen, P. et al. Chem.-Biol. Interact. 209, 14-24 (2014).
For Research Use Only. Not for use in diagnostic procedures.
>> Update My Profile | Subscribe to Access Agilent | Article Directory