Access Agilent eNewsletter, June 2014
>> Update My Profile | Subscribe to Access Agilent | Article Directory
Lipidomics: new tools and applications start here
By Nigel Skinner
Agilent Segment Marketing Manager, Life Science Research
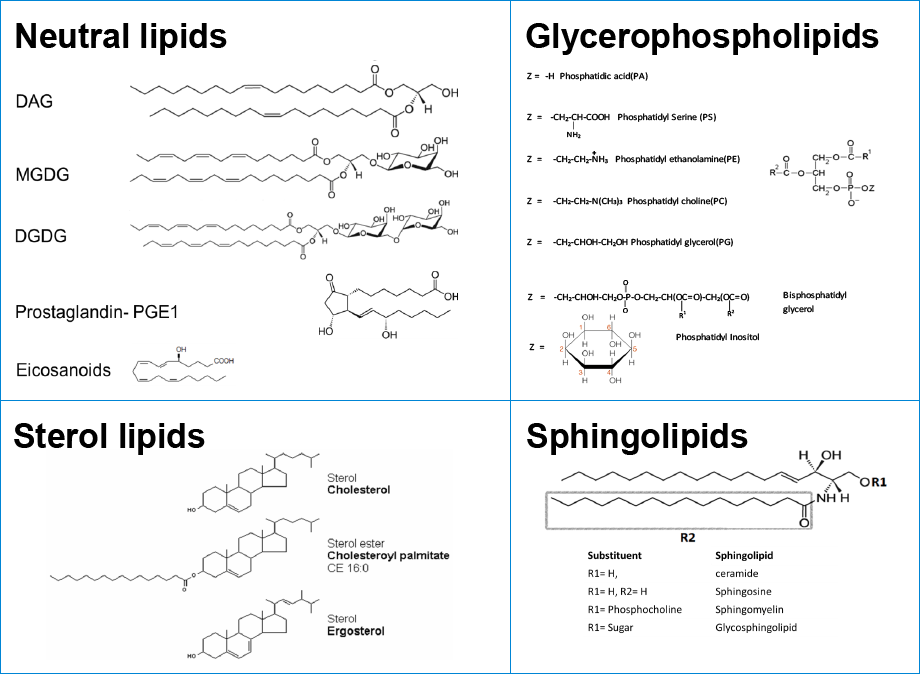
Lipidomics is the systematic study of the entire lipid constituents of a cell/tissue/organ/organism. Mass spectrometry (MS) is one of the most widely recognized technologies in lipidomics research today. It is used for the identification and quantitation of thousands of lipid molecules that constitute the full lipid complement of a biological system, called the lipidome.
 Enlarge
Enlarge
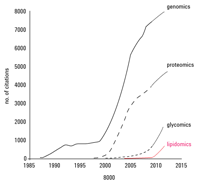
Figure 1. Lipidomics is an emerging field.
Lipids are generated and metabolized by enzymes that are influenced by the environment of a given biological system – for instance, by diet and temperature. Some estimates have placed the number of distinct chemical entities within the lipidome between 10,000 and 100,000. Today there is an increasing awareness across many disciplines of the critical importance of lipids in all aspects of life, such as energy homeostasis, membrane structure and dynamics, and signaling. Because imbalances in lipid metabolism contribute to diverse phenotypes and disease states, there are an expanding number of drugs that target lipid metabolic and signaling pathways. Examples include cholesterol-lowering agents (statins) and cyclo-oxygenase inhibitors. Drug discovery researchers are especially interested in identifying specific regulators of numerous targets – including phosphatidylinositol 3-kinases, nuclear hormone receptors, sphingosine, and ceramide kinases – in disease research from cancer to metabolic disorders. Currently, many biologically important lipids can be analyzed on a fairly routine basis, though lipidomics is still very much an emerging field, compared to genomics and proteomics (Figure 1).
Understanding discovery lipidomics
Compared to proteins, the MS of lipids is very different due to their large chemical diversity [1], making it difficult to comprehensively measure the lipidome of a cell or tissue in a single experiment. ‘Discovery’ lipidomics has become a popular approach. This involves the examination of an untargeted suite of lipids to find the ones with statistically significant variations in abundance within a set of experimental versus control data samples. Their chemical structure is then determined. Such methods should have high mass accuracy and resolution, a characteristic of time-of-flight MS. Fragmentation of an ion of interest is then used for identification. Analysis of fragmentation pathways has led to a detailed understanding of ‘bonding’ between the different building blocks in lipids (such as fatty acids, sphingoid bases, and head groups). It has also formed a basis for ‘shotgun’ lipidomics in which precursor lipids are determined based on characteristic fragment ions [2].
Lipids and their metabolites integrate many cellular functions. Thus energy homeostasis is tightly coupled to fatty acid metabolism, and fatty acids are key building blocks of many cellular lipids. Given that cancer cells are dependent on fatty acids for the synthesis of membranes and signaling lipids, fatty acid synthase is considered a potential therapeutic target. This known lipid ‘signaling’ network is poised for a great expansion, particularly for human disease [3].
Meaningful answers in lipid discovery experiments are dependent not only on MS, but also on specialized data analysis software that facilitates cheminformatics, bioinformatics, and statistical analyses and provides a search program using an extensive lipid database. The Agilent Lipidomics Workflow Guide [4] walks you through using our Agilent Mass Profiler Professional Software products, as well as a non-Agilent lipid database program. This approach delivers meaningful answers to research questions more efficiently than using typical mass spectral data analysis and identification programs.
From discovery to insight
Typical analytical readouts in ‘discovery’ approaches include retention time, mass-to-charge ratio, and information on fragment ions. ‘Targeted’ analysis delivers a matrix of lipid identities and their intensities. Typical informatic frameworks can include data processing (peak integration, identification, and normalization), statistics, and integration into pathways such as KEGG (Kyoto Encyclopedia of Genes and Genomes). Pathway analysis lets you connect the lipid with a biological process or condition. SimLipid, a product of PREMIER Biosoft International, is a lipid search program that rapidly searches an extensive lipid database. Agilent and PREMIER Biosoft have worked together to integrate the lipid identification at MS and MS/MS levels using SimLipid software in Agilent’s unique Compound Exchange Format. This allows seamless portability of the lipid output data between different Agilent programs and SimLipid.
Next frontier: ion-mobility mass spectrometry
Ion-mobility mass spectrometry (IMS), which combines information on molecular shape with mass/charge, has not yet been extensively applied to the analysis of lipids. Biophysical studies have shown that the double bonds of fatty acyls determine the conformation of lipids in bilayers, and this structural characteristic might also affect the collisional cross-section. It is also conceivable that ion mobility is affected by head-group geometry (which is impacted by phosphorylation and glycosylation). Thus, it is likely that IMS will provide valuable information that is otherwise difficult to obtain.
Resources and solutions for integrated biology
Agilent is committed to integrated multi-omics discovery and integrated biology, with a comprehensive portfolio of sample prep, LC columns, and chromatography supplies. Recent advancements include the latest version of our GeneSpring bioinformatics software, designed for pathway-centric multi-omic data integration. Take a moment to explore Agilent Integrated Biology solutions and metabolomic resources today.
References
- E. Fahy et al. J. Lipid Res. 46, 839 (2005).
- M. R. Wenk. Nat. Rev. Drug Discov. 4, 594 (2005).
- M. P. Wymann R. Schneiter. Nat. Rev. Mol. Cell Biol. 9, 162 (2008).
- Anon. Agilent Lipidomics Workflow Overview. Agilent Technologies, Inc. Publication number 5991-1644EN (2012).
For Research Use Only. Not for use in diagnostic procedures.
>> Update My Profile | Subscribe to Access Agilent | Article Directory