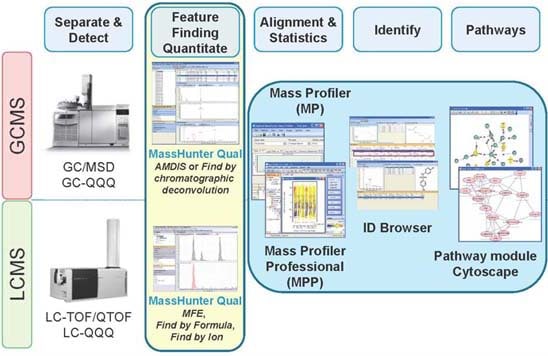
Metabolomics at Agilent: Driving Value with Comprehensive Solutions
|
|
By Darlene J.S. Solomon, Ph.D., Agilent Chief Technology Officer, and
Agilent is the leader in solutions for metabolomics with the most comprehensive product portfolio that spans the metabolomics workflow. Metabolomics is one of the fastest growing areas of life sciences research. It is used in plant research for crop improvement, clinical research to better understand diseases, nutritional research to improve health through diet, pharmaceutical research to identify biomarkers for disease and spot potential toxic side effects of drug candidates earlier in development, and fermentation research to bioengineer organisms to produce valuable products. In this article, we will share our perspectives on the following topics:
The number of primary metabolites needed to explore a person’s health comprehensively is orders of magnitude lower than the number of genes, RNA transcripts or proteins required for similar study. Estimates of primary metabolites range from 2,500 to 6,000, though these numbers are much higher for secondary metabolites. Genomics involves the study of about 25,000 genes, transcriptomics about 100,000 RNA transcripts, and proteomics about 1,000,000 proteins, including post-translational modifications. Modern metabolomic research has its origins in nuclear magnetic resonance (NMR) spectroscopy for the study of the chemical structure of metabolites. Jeremy K. Nicholson of the Imperial College London has used NMR since the early 1980s to study, for example urine from animals that had been poisoned by different drugs. When he submitted a paper to a scientific journal in 1987, the editor said that this research had no interest to anyone. That was 10 years in advance of the first paper that would really call itself metabolomics.(1) Since the late 1990s, mass spectrometry-based studies of metabolomics have become more prevalent than NMR. Researchers realized they could use gas chromatography/mass spectrometry (GC/MS) to study complex samples and then use statistical tools to extract information from chemical systems. Later, researchers expanded into liquid chromatography/mass spectrometry (LC/MS), driven by the advent of affordable, accurate mass, time-of-flight (TOF) instruments. Metabolomics research applications Basic metabolomics research: Medical schools, research hospitals, universities and academic institutes conduct basic research in metabolomics to understand human diseases and biological processes. For example, researchers at the Sarah W. Stedman Nutrition and Metabolism Center are studying the complexity of type 2 diabetes using key metabolomic methodologies, including NMR and mass spectrometry, to understand the molecular pathways that contribute to type 2 diabetes. They’re integrating analysis of genomic, transcriptomic and metabolomic datasets to learn more about metabolic regulatory networks and diabetes mechanisms. (2) Metabolomics for biofuels production: Amyris has demonstrated that microbial metabolism can convert biomass to fuels cost-effectively – as well as to chemicals and pharmaceuticals. By altering the metabolic pathways of microorganisms, such as yeast, these genetically modified microorganisms serve as living factories that produce molecules for use as chemicals and transportation fuels. In 2009 the U.S. Environmental Protection Agency officially registered Amyris’s plant derived diesel fuel, making it the first time a hydrocarbon-based fuel made from fermentation of plant material was registered for commercial use. (4) Fueled by research, Agilent Technologies has long been a leading supplier of gas chromatographs, liquid chromatographs and mass spectrometers known for excellent performance, reliability, ruggedness and ease of use. Agilent continues to invest in software and informatics tools to enable customers to achieve new levels of insight from their experiments.
Challenges are significant in the separation and detection of metabolites following sample preparation:
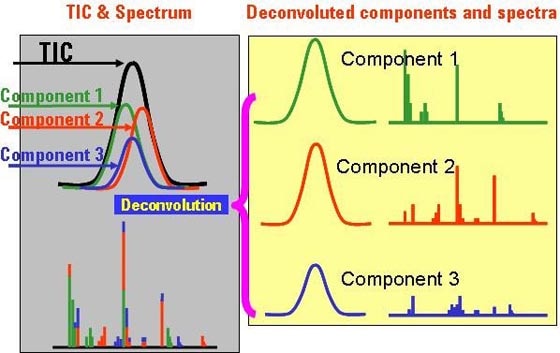
Agilent’s GC/MS and LC/MS can both be used to separate and detect metabolites, and each technique has its own strengths. Most researchers find they need both GC/MS and LC/MS to get a comprehensive metabolite picture because of the vast chemical diversity of metabolites and their wide variation in abundance. Discovery metabolomics Agilent's GC/MS Metabolomics Profiling and Identification Solution couples the Agilent 7890A GC with the 5975 GC/MSD to combine performance, reliability and cost-effectiveness for discovery and routine profiling studies. These systems also enable rapid, confident sample identification through searching of electron ionization (EI) spectral libraries. The 7000A Triple Quadrupole GC/MS system also may be used for discovery metabolomics and offers more sensitivity and specificity for targeted metabolomics. By integrating Agilent instruments, columns and software, researchers can shorten their path to meaningful results. Key attributes of TOF and Q-TOF instruments are accurate mass, acquisition speed, mass resolution and dynamic range. Agilent’s TOF and Q-TOF accurate mass capability contributes significantly to metabolomics research. When researchers measure a complex sample, the resolution and mass accuracy possible with Agilent’s TOF technology give better measurements to yield more information from the sample. Mass accuracy and resolution reduce the importance of chromatographic analysis because the mass spectrometer does more of the work. Targeted metabolomics Both the 5975C single quad GC/MS and 7000A triple quadrupole GC/MS system can be used for quantitation. The 5975C single quad GC/MS is the more cost-effective option; however, where the very lowest levels of detection and quantitation are required for extremely low abundance metabolites, Agilent's 7000A triple quadrupole GC/MS system combines unmatched sensitivity and selectivity with everyday reliability and ease of use. If customers require one system for discovery and targeted metabolomics, they can use the versatile 7000A system with no compromises. Software challenges In 2009, results of a survey conducted by American Society For Mass Spectrometry (5) indicated that 81 percent of respondents attributed metabolomics challenges to software, not hardware. This percentage included compound identification (35%), biological significance (27%), data processing (14%) and statistical analysis (5%). Agilent’s software suite includes capabilities that do much of the “heavy lifting” for researchers in data analysis and compound identification. Software enables peak finding to locate and identify the metabolites needed for biological understanding. Software also is required to normalize and correct the data, and analyze the data statistically to find meaningful differences in sample sets. Agilent’s Mass Hunter Qualitative software processes data and finds features. Mass Hunter Qualitative and MassHunter Quantitative have versions for GC/MS and LC/MS that are conceptually similar but work differently based on the way the ions are made. GC/MS typically uses electron impact to ionize compounds, and the result is a set of spectral ions that correspond to a compound. LC/MS typically utilizes electrospray ionization that results in molecular ions with almost no fragments. Feature finding/compound identification GC/MS - Agilent Deconvolution Reporting Software (DRS) enables separation and identification of two or more chemical compounds that emerge from a chromatographic column and flow into the mass spectrometer with incomplete chromatographic separation. In a complex matrix, a chromatographic peak may be a composite of overlapping components. Agilent’s DRS uses a mathematical technique that separates overlapping mass spectra into “cleaned” spectra of the individual components. The simplified total ion chromatogram (TIC) below illustrates the process:
LC/MS - Molecular feature extractor: The Agilent MassHunter molecular feature extraction software is designed to rapidly find peaks in the total ion chromatogram produced when a sample was analyzed using an Agilent LC/MS accurate mass TOF system. The software removes the chemical background from the three dimensional LC/MS dataset, finds the true ion signals, groups the chemically related ion signals (isotopes, adducts and dimers) and displays peaks in a compound chromatogram with associated pure spectra. The result is a compound table with associated chromatograms and pure spectra with each compound being quantified. The result of this data process is ready for further analysis. For LC/MS-based metabolomics, Agilent has partnered with Gary Siuzdak, Ph.D. to commercialize the METLIN Metabolite Database, a repository of useful metabolite information. As part of the partnership a LC/MS/MS metabolite library is being produced for Agilent Q-TOF instruments. This library will allow for the identification of metabolites by spectral pattern matching. Metabolomics is such a new science that typically only 20 percent of metabolites can be identified by database searching or MS/MS library matching. Molecular formula generator: For those metabolites that could not be identified by the use of the METLIN database and MS/MS library,researchers can derive molecular formulas for the unknowns based on TOF and Q-TOF mass spectral data. Agilent’s MassHunter molecular formula generator takes full advantage of the mass accuracy of the data, and then uses additional mass spectral information (isotope ratios and isotope mass values) to logically narrow the list of possible formulas. It delivers a list of candidate molecular formulas and the relative probability that each formula is the correct one. This process significantly reduces data interpretation time and increases the value of accurate-mass analysis. Statistics and biological pathway analysis Mass Profile Professional is designed specifically for MS data to move results into a statistical software package for analysis. Although third-party statistical packages are available, Agilent’s statistical package is designed for mass spectral data. This has many benefits to the researcher such as the ability to query the data as mass spectral data to find, for example, all the compounds that are in the data set that have the fragment ion 73. Biological meaning can only come if compounds are identified, so growing Agilent’s metabolite libraries is critical to advancing metabolomics. Agilent continues to work with leaders in the field to co-develop libraries that will help further research and understanding. This commitment to improve compound identification is tied to Agilent’s interest in pathway analysis as only identified metabolites can be used to analyze biochemical pathways. Challenges to understanding the role of metabolites involve relating their functions to surrounding compounds and observing interactions in biological pathways. To enable and facilitate biological interpretation, Agilent built a pathway analysis tool into Mass Profiler Professional that facilitates interpretation with known pathways. With these tools researchers also can build their own pathways with observed data and connect common chemical names with biological synonyms found in biological pathways. Mass Profiler Professional data is easily interchanged and correlated with data from other products in the Agilent GeneSpring Analysis platform, providing valuable biological insight at the intersection of proteomics, metabolomics, genomics and biomarker research. Orthogonal detection and molecular structure identification With the recent acquisition of Varian, Agilent’s portfolio now includes NMR spectroscopy with NMR products spanning the entire workflow, from sample preparation to software for data acquisition and processing. Varian produced the first commercial NMR more than 50 years ago and continued its technical advances to stay at the forefront of this broadly applicable technology. In metabolomics, NMR analysis provides a powerful compliment to mass spectrometry based methods. Researchers can use NMR to directly investigate many types of samples from whole tissues to cell cultures to biological fluids without significant sample preparation or adulteration. Localized NMR spectroscopy can be used to do in vivo investigation of spatially segregated structures within an intact organism. The NMR experiment is non-destructive and inherently quantitative due to unit response factors for all molecules. Because the spectroscopic property that NMR interrogates is unrelated to those used in chromatography or mass spectrometry, NMR provides a completely orthogonal detection mechanism. While there are benefits to the direct analysis of an intact metabolic sample, the ability to differentiate responses from individual components is a challenge because these samples are typically complex mixtures. The Premium COMPACT ultra high-field NMR systems provide the dispersion needed to resolve individual components. When coupled with Agilent’s proprietary cold probe technology, these systems deliver the performance required to detect high-picogram quantities of metabolites in a sample. Metabolomics studies using NMR entail a very broad range of sample characteristics so the software used to control the spectrometer must be flexible enough to accommodate this diversity. The NMR community knows Agilent’s VnmrJ software as the only spectrometer control package featuring open architecture that users can customize easily. The synergy between NMR and MS to clarify unknown structures has been recognized for many years. NMR readily discriminates positional or structural isomers and, in most cases, provides the information required for de novo characterization of chemical structures. Agilent now seamlessly provides all the instrumentation required to detect, isolate and analyze the chemical structure of any component in a complex metabolic mixture. NMR system configuration for optimized metabolomics research presents many challenges, including the following:
The technology and experience that Varian brings allow Agilent to deliver automated acquisition of high-quality NMR data for metabolomics research. As is the case for LC/MS and GC/MS, however, NMR data reduction remains the time intensive part of the metabolomics study. Several available tools can help to reduce this burden, and focus in the future will be to make contributions to this area. The chart below provides a brief look at the benefits and tradeoffs of selecting GC/MS, LC/MS or NMR for metabolomics.
Synergies across Agilent At the foundation of Agilent solutions for metabolomics are significant and unique synergies that enable high performance and low cost. The ability of Agilent Research Laboratories to see across all businesses in the company enables Labs’ researchers to discover how innovations in one area could contribute to one or more very different ones. QTOF and QQQ: Agilent’s QTOF and triple quad analyzer platforms work well together because of leveraged technology across both platforms: same quadrupole, same common collision cells and other components that enable excellent reproducibility for integrating data across systems. With a common collision cell fragmentation patterns on the QTOF are similar to fragmentation patterns on the QQQ, a desirable outcome that is the result of thinking through customer needs from discovery through targeted analysis. TOF and oscilloscopes: Agilent leverages high-speed oscilloscope technology to TOF capability, because digitization in both areas involves high-speed data conversion and digital signal processing. At low mass, ions arrive so rapidly at the detector that high speed digitization is required to capture the arrival rates. Agilent is the only MS vendor with proprietary data conversion technology and therefore not limited to data converters available commercially. By leveraging multiplexing developed from the oscilloscope teams, Agilent can use a high-gain channel to measure a weak signal, a low gain channel to measure a strong signal and still get mass accuracy for both. Multiplexing extends the dynamic range of the TOF instrument to see the lower-intensity and high-intensity signals present in metabolomics study. World-class development and manufacturing: Agilent makes ongoing investments to improve performance, drive down cost and make LC, GC and MS affordable, practical and widely accessible. Although leading edge technologies are initially expensive, Agilent works to understand and engineer system components so they’re both high performance and relatively low cost. Agilent’s ability to scale and leverage its worldwide manufacturing capability has resulted in a leading position of expertise and efficiency while maintaining a reputation for quality and reliability. Manufacturing teams around the world work together to leverage processe and share best practices, and are linked closely with R&D to build high-quality, reliable instruments. Because Agilent’s GCs today are much less expensive compared to their cost two decades ago, they’re used more broadly around the world. The more researchers who can afford to work on metabolomics, the faster science and understanding will progress. Agilent is advancing metabolomics R&D with what are referred to as win-win-win innovations. Customers win with access to new technology to investigate questions they couldn’t address or answer before. Science wins because Agilent solutions enable better understanding of biology and the world, and Agilent wins because we recoup the initial investments to continue innovating future measurement solutions that advance the leading edge. Agilent also continues to collaborate with scientists and thought leaders to develop innovative solutions to problems of the next year and the next decade. Examples of these valued partnerships include the following:
We are charting future directions and supporting the entire innovation process to get inventions out of our laboratories and into products and solutions that create real value for our customers. # # # References Nicholson has continued to use NMR as a tool for metabolomic research to study disease and other pathological processes. NMR is a nondestructive technique that requires minimal sample preparation, and although NMR sensitivity is lower than mass spectrometry, it enables observation of many metabolites, especially at high magnetic field strengths. (Evaluation of Full-Resolution J-Resolved 1H NMR Projections of Biofluids for Metabonomics Information Retrieval and Biomarker Identification, Anal. Chem., 2010, 82 (5), pp 1811–182. 3. Boudonck KJ, Mitchell MW, Német L, Keresztes L, Nyska A, Shinar D, Rosenstock M.“Discovery of Metabolomics Biomarkers for Early Detection of Nephrotoxicity.” Toxicol Pathol. (37)3 2009: 280-92. 4. Newman, Jack. “Microbial Fermentations for Diesel Fuel Production: The Role of Metabolite Analysis in Strain Improvement.” Metabolomics 2010 Conference Program, June – July, 2010: pg. 34. |
||||||||||||||||||||